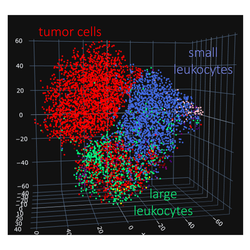
Whether cells, nerves, or blood vessels, fully capturing the morphology of biological structures is a challenging task. We develop methods to represent meaningful visual features captured in images in a label-free manner. In particular in biomedical images where labeling requires often additional expertise, the possibility to explore image data in an unsupervised manner can guide further targeted analyses.
Highly differentiated diseases require large-volume, high-resolution datasets and novel analytical approaches to find recurring patterns in large spectrum of genetic and molecular variation. In collaboration with medical and experimental groups at the Charité University Hospital Berlin we address the question of high molecular heterogeneity in Triple Negative Breast Cancer (TNBC). This most aggressive breast cancer subtype with high rates of recurrence and mortality offers currently no therapy options.
In this project we expand over the existing knowledge of the genetic heterogeneity between and within TNBC tumors, by investigating the differences in their transcriptome, proteome and morphology as well as the extracellular matrix components and immune cell environment. Our analysis is based on patient samples from prospective clinical trials at multiple levels.
We strive to establish a novel type of biomarkers that instead of individual molecule levels will integrate multiple types of information and will be quantified with the use of machine learning and mathematical models.
C. elegans, a tiny nematode worm, is used to study a broad range of questions in biology, from diseases to neural function. This apparently simple organism shows a broad repertoire of behaviors incomprehensible to human observer. These behaviors might be representative of its health and disease phenotype.
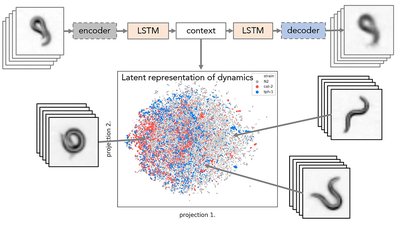
We employ deep learning methods to quantify and search for distinct motion patterns representative of worm molecular phenotype. The grand challenge of this project involves finding ways to represent worm posture and dynamics. Inspired by methods for image and language processing we aim to find meaningful representations of words and sentences in the language of worm behavior

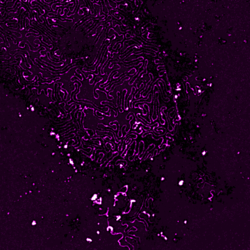
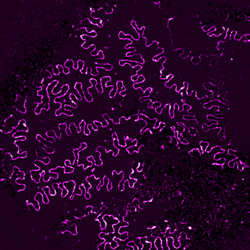
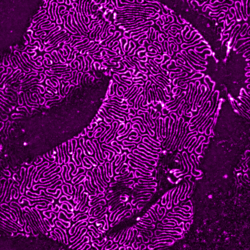
In collaboration with the nephrology lab at the University Hospital Cologne, we explore super-resolution microscopy images of molecular morphology of kidney. The site of kidney filtration barrier is composed of specialized cells called podocytes which extend into formation termed foot processes (FPs). These formations stick to neighboring podocytes in an interdigitating pattern. The slits between adjacent FPs allow the passage of fluid and solutes, and they are spanned by a specialized cell-cell contact between FPs. Key proteins for the slit function – Podocin and Nephrin – can be imaged with super-resolution microscopy allowing to view the intricate shapes of the filtration barrier while a mouse model of human chronic kidney disease allows to precisely follow its progression.
In this project we adopt machine learning approach to comprehensively analyze and quantify morphological changes of podocytes related to the kidney filtration decline. FPs form intricate patterns difficult to describe manually or with a set of hand-designed visual parameters. Not only size and shape of individual FPs might play a role in filtration but also their collective configurations within the tissue. Although differences between healthy and diseased adult tissues are striking, these differences are less apparent between young, pre-symptomatic individuals. Here we combine machine learning methods with super-resolution microscopy and the unique mouse model to search for morphological markers of the disease.



We explore image encoding and classification models to perform predictions on various types of tumor histopathology images. The questions we are interested in is tumor detection, tumor type classification, as well as prediction of therapy response. Beyond mere classification, we use attention mechanism to gain insights into the visual information that is key for a given classification task. Based on segmentation and staining we aim to provide interpretable cues on which cell types, their spatial distributions and cell-cell interaction might be specific to certain tumor types or predictive of patient clinical outcomes.